Targets
Target lists are collections of peak definitions used to extract MS intensities for specific metabolites. The Targets tab allows you to manage, review, and refine these definitions before processing. To import a target list, click the LOAD TARGETS button. This opens a file browser where you can navigate your filesystem and select a CSV file containing your peak definitions.
Tip: Click the help icon (small "i" symbol) next to the "Targets" title to take a guided tour of this section.
The Targets Table
The target list can be provided as a CSV file with the following columns. A MINT-compatible template can be downloaded in the Targets tab (DOWNLOAD TEMPLATE button on the top right corner).
Targets table columns and descriptions
| Column Name | Description |
|---|---|
peak_label |
Unique metabolite/feature name |
peak_selection |
True if selected for analysis |
bookmark |
True if bookmarked |
mz_mean |
Mean m/z (centroid) |
mz_width |
m/z window or tolerance |
mz |
Precursor m/z (MS2) |
rt |
Retention time (default: in seconds) |
rt_min |
Lower RT bound (default: in seconds) |
rt_max |
Upper RT bound (default: in seconds) |
rt_unit |
RT unit (e.g. s or min; default: in seconds) |
intensity_threshold |
Intensity cutoff (anything lower than this value is considered zero) |
polarity |
Polarity (Positive or Negative) |
filterLine |
Filter ID for MS2 scans |
ms_type |
ms1 or ms2 |
category |
Category |
formula |
Chemical formula (used to derive m/z when mz_mean is missing) |
maven_id |
Maven ID or Group ID (optional identifier) |
adduct_name |
Adduct name (e.g. [M+H]+); optional and can be auto-populated |
score |
Optional legacy score field |
notes |
Free-form notes |
source |
Data source or file |
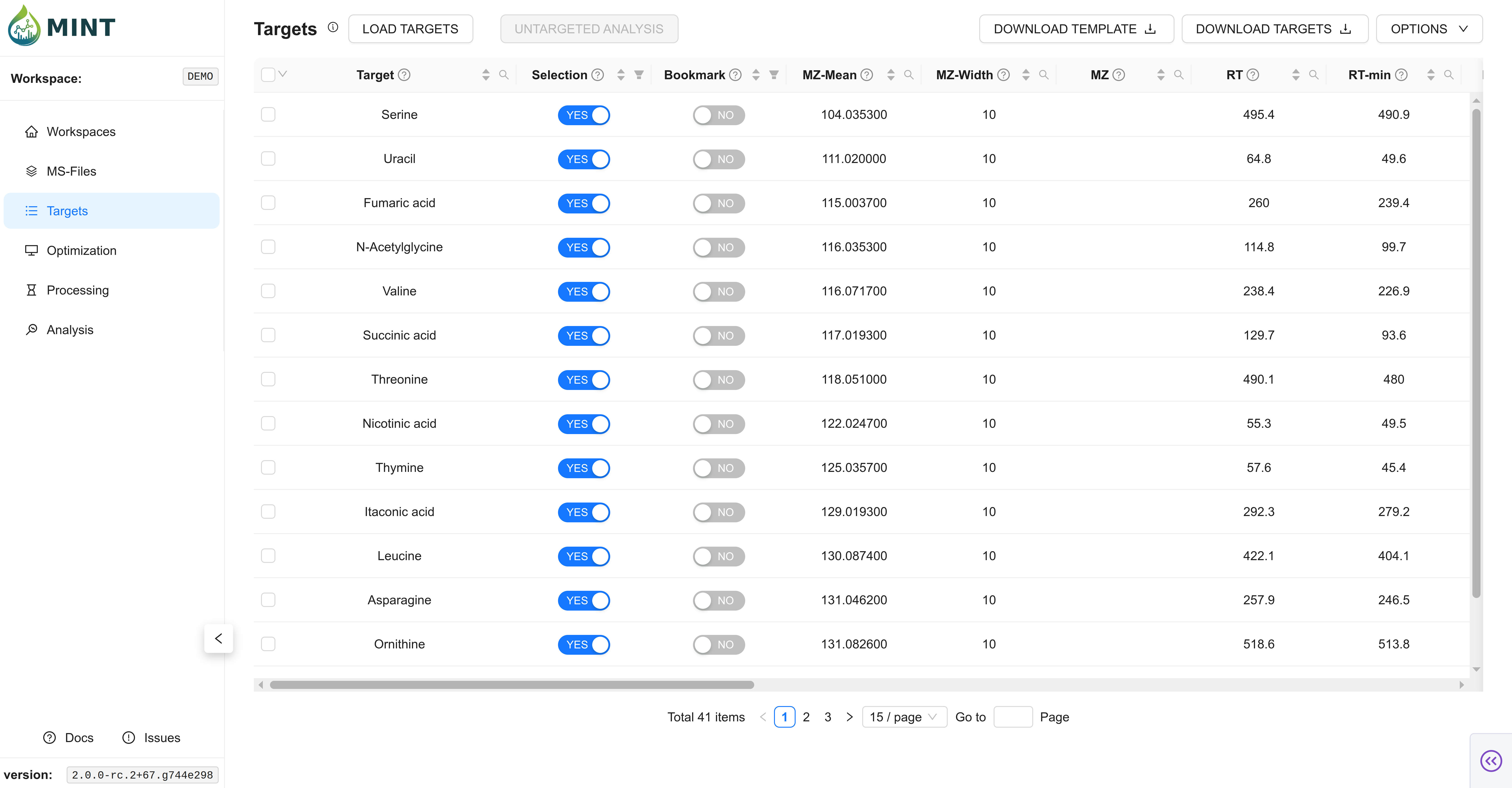
Once loaded, your targets are displayed in an interactive table with the following key columns:
- Peak Label: The unique identifier for the metabolite or feature.
- Selection & Bookmark: Selection is meant for selection of targets for processing. Bookmarking is meant for bookmarking specific targets for later use.
- MZ Data:
mz_meanandmz_widthdefine the mass-to-charge ratio window for extraction. - Retention Time:
rt_minandrt_maxdefine the expected time window for the peak. Thertcolumn typically represents the expected retention time of the peak. - Filtering: Each column header includes a filter icon, allowing you to search for specific compounds or filter by values.
Selection and Bookmark Semantics
MINT uses peak_selection and bookmark with a few workflow rules that affect Optimization and Processing:
- Selection drives default target scope: selected targets are included first.
- If no targets are selected, MINT falls back to treating this as all targets selected (instead of processing zero targets).
- Bookmark implies selection: when a target is bookmarked, MINT also marks it selected so bookmarked subsets remain processable.
- Bookmarked-only mode (Processing): enabling
Bookmarked Targets Onlyintersects the processing scope with bookmarked targets.
Importing from External Software
MINT supports importing target lists directly from other software formats, such as EL-MAVEN (peak lists and group exports).
Key features of this import functionality include:
- Format Support: Reads CSV, TSV, Excel, and JSON (EL-MAVEN group exports) files.
- Automatic Column Mapping: MINT automatically detects and renames columns from common formats to the MINT standard.
compound,compoundName,name->peak_labelmeanMz,medMz,parent,row m/z->mz_meanmeanRt,medRt,expectedRt,row retention time->rtformula,parentformula->formula
- Unit Conversion: Retention times in EL-MAVEN files (typically in minutes) are automatically converted to seconds.
- Priority Handling: If a file contains multiple potential columns for the same value (e.g., both
meanMzandmedMz), MINT automatically selects the most appropriate one based on a predefined priority list.
Untargeted analysis (via Asari)
If you don't have a pre-defined target list, MINT can automatically detect features in your processed files using Asari.
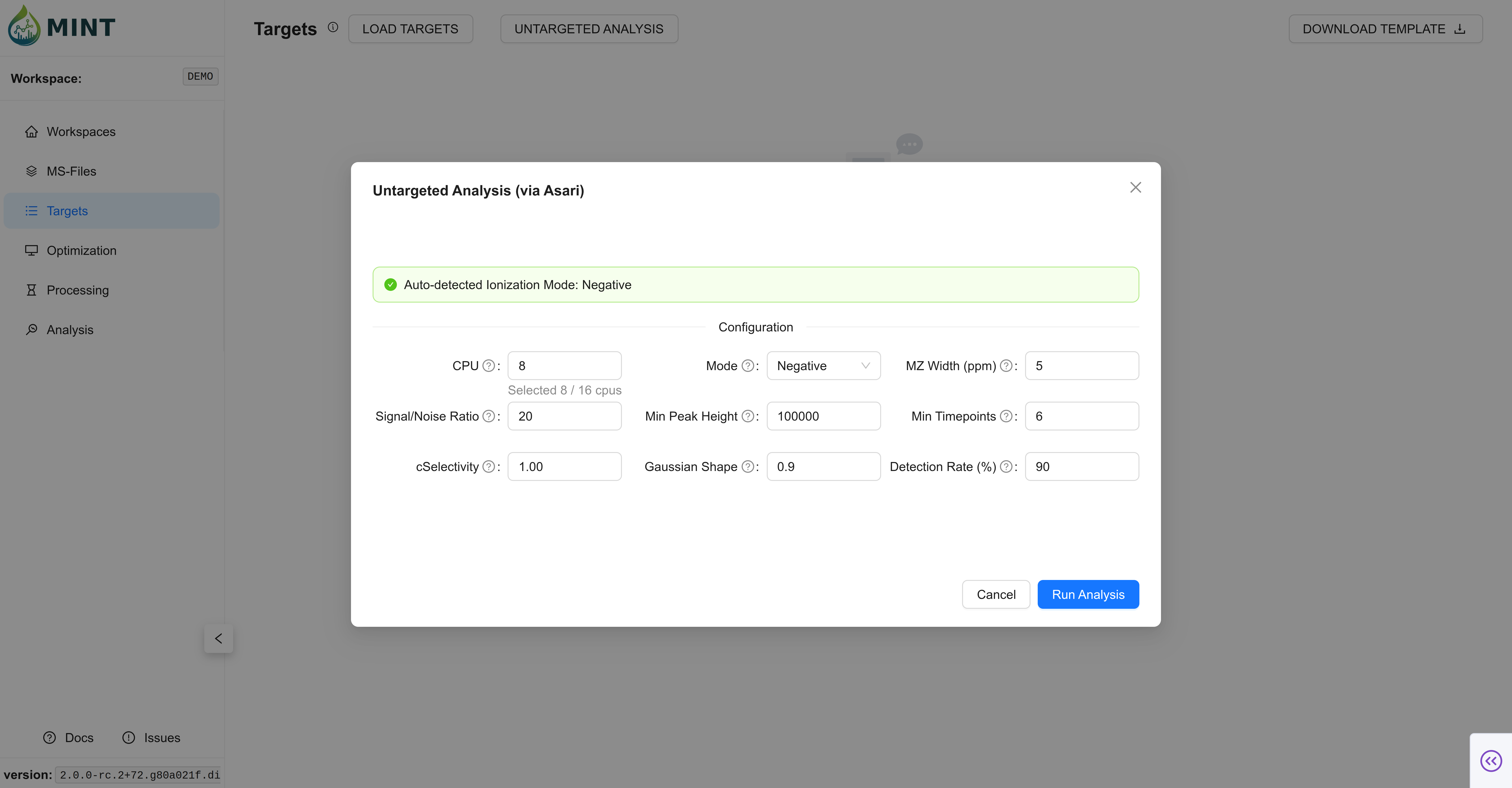
- Click the
Untargeted analysisbutton to open the configuration modal. - Adjust the parameters to suit your data (ionization mode, mass tolerance, signal-to-noise ratio, etc.).
- Click
Run Analysis. MINT will process the files in the background and populate the table with detected features.
Key Parameters:
- CPU: Number of cores to use for parallel processing.
- Mode: Ionization mode (
PositiveorNegative). - MZ Width (ppm): Mass tolerance for grouping peaks.
- Signal/Noise Ratio: Minimum signal-to-noise ratio for peak detection.
- Min Peak Height: Minimum intensity required for a peak to be considered.
- Min Timepoints: Minimum number of scans required to define a peak shape.
Import Validation and Auto-Fill
MINT only strictly requires peak_label plus enough RT information to define a window. For each target row, MINT accepts valid RT input patterns and auto-fills missing values. In MINT terminology, those bounds define the target Region of Interest (ROI). Below, we explain in detail some of the rules MINT uses to fill missing elements in the targets table. Click the banner to expand.
RT Auto-Derivation and Validation
RT Auto-Derivation and Validation
Before MINT derives any missing RT fields, it first normalizes RT values to seconds. This matters when rt_unit = min: for example, rt = 2.0, rt_unit = min is converted to 120 s first, and only then does MINT derive bootstrap ROI bounds around that value.
rt_min+rt_maxonly: MINT derivesrtas the midpoint.rtonly: MINT initializesrt_minandrt_maxusing a temporary symmetric bootstrap window (rt ± 5.0 s) so targets can be processed immediately. The±5.0 srule is an initial import fallback, not the final recommended span. After chromatograms are computed, refine RT bounds in the Optimization tab using the observed peak shape and apex.rt+rt_min: MINT derivesrt_maxsymmetrically aroundrt.rt+rt_max: MINT derivesrt_minsymmetrically aroundrt.- All three (
rt,rt_min,rt_max): MINT keeps them, then validates consistency.
Invalid RT inputs:
- Only
rt_min(withoutrtorrt_max) - Only
rt_max(withoutrtorrt_min) - Missing all RT fields (
rt,rt_min, andrt_max)
Additional RT validation done during import:
rt_minmust be less thanrt_maxrt_min/rt_max/rtcannot be negative- If
rtis outside[rt_min, rt_max], MINT resetsrtto the span midpoint rt_unitvalues in minutes are converted to seconds before derivation/validation, and stored as seconds
Defaults and Overrides
Defaults and Overrides
During import, MINT applies a few defaults and override rules so target rows are immediately usable:
mz_widthdefault: Ifmz_widthis missing, MINT sets it to10.0(ppm).peak_selectiondefault: If missing/blank, targets default toTrue(selected).bookmarkdefault: If missing/blank, targets default toFalse.ms_typesource of truth: MINT derivesms_typefromfilterLine:filterLinepresent ->ms2filterLineabsent ->ms1
ms_typeconflict handling: If your file providesms_typethat contradictsfilterLine, MINT keeps the derived value and logs a warning.rt_unitnormalization: Units in minutes are converted to seconds before RT derivation, and stored withrt_unit = s.
Duplicate Label Handling
Duplicate Label Handling
When imported targets contain repeated peak_label values, MINT resolves them automatically:
- Duplicate labels are renamed to
peak_label@RT(for example,Succinate@315.20) when RT is available. - Exact duplicate rows after renaming (same final label) are collapsed to one entry.
- A summary of duplicate-label handling is reported in processing notifications/logs.
Smart Enrichment After Import
Smart Enrichment After Import
After parsing and validation, MINT applies additional enrichment to improve incomplete target tables:
- Polarity inference from loaded MS files: If loaded samples have one consistent polarity, missing target polarities are auto-filled from that value.
- Polarity mismatch warning: If a target explicitly specifies a different polarity than the loaded MS files, MINT keeps the row but emits a warning.
- Adduct auto-fill: If
adduct_nameis missing, MINT derives defaults from polarity ([M+H]+for positive,[M-H]-for negative). - Formula-based
mz_meanfill: Ifmz_meanis missing andformulais present, MINT attempts to calculatemz_meanfrom formula + polarity. - MS1 precursor cleanup: For
ms1targets,mz(precursor field) is cleared to avoid redundant/confusing values.
Backup Snapshot
Backup Snapshot
Whenever targets are imported (file upload) or generated via Asari, MINT writes a workspace-local backup snapshot:
- Path:
<workspace>/data/targets_backup.csv - Purpose: Fast recovery/export of the current target table and safer downstream workflows.
- Update behavior: The file is refreshed after each successful target load/generation.
Options Menu
The Options dropdown (top right) provides bulk actions for list management:
- Delete selected: Remove currently checked rows from the workspace.
- Clear table: Remove all targets.