Analysis
The Analysis tab is where you explore processed results interactively across multiple statistical and quality-control views.
Tip: Click the help icon (small "i" symbol) next to the "Analysis" title to take a guided tour of this section.
Global Controls
The top toolbar controls the data feeding all Analysis views.
- Metric:
Peak Area,Peak Area (Top 3),Peak Max,Peak Mean,Peak Median,Peak Area (EMG Fitted), andConcentration(when SCALiR output exists). - Transformations: Apply statistical transformations to the raw data:
- None (raw): Use raw values.
- Z-score: Standardize features (mean=0, std=1).
- Rocke-Durbin: Variance stabilization.
- Z-score + Rocke-Durbin: Standardize, then stabilize variance.
- Group by: Choose grouping by
sample_typeor metadata group columns (group_1...group_5).
Data scope and defaults
- Analysis uses samples flagged
use_for_analysis = TRUE. - Default normalization is view-specific:
PCA,QC,Violin,Bar,Comparison:Rocke-Durbint-SNE,Clustermap:Z-score
- If results are missing, Analysis prompts you to run Processing first.
- Missing group labels are treated as an explicit
"(... unset)"group and shown with neutral color.
Analysis Views
The left sidebar in the Analysis tab allows the user to switch between different views:
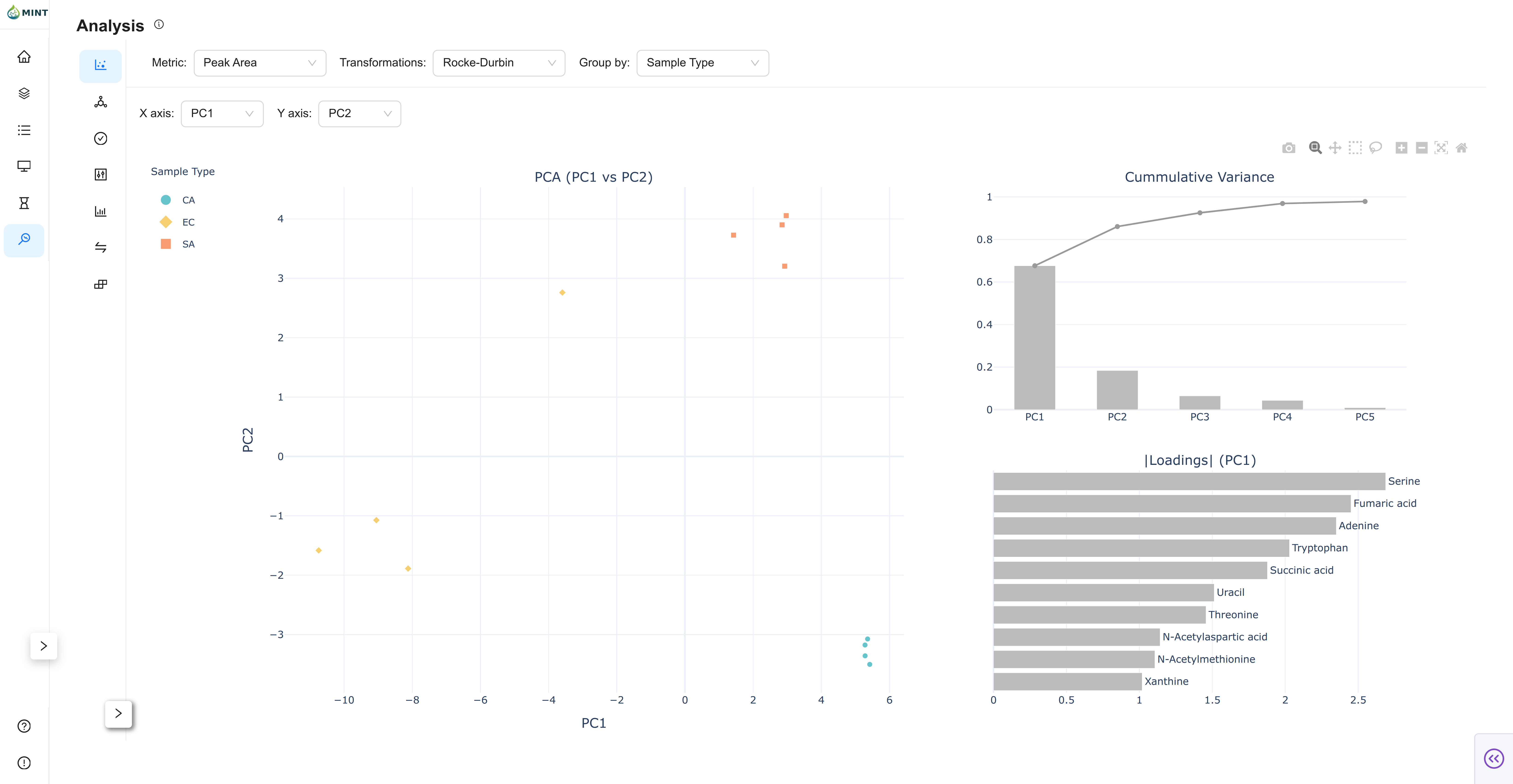
PCA View
PCA projects samples to principal components and highlights structure across groups.
- Axis controls: Choose X/Y among
PC1...PC5. - Score plot: Sample projection colored by the selected grouping field.
- Variance panel: Per-component and cumulative explained variance.
- Loadings panel: Top absolute loadings for the active component.
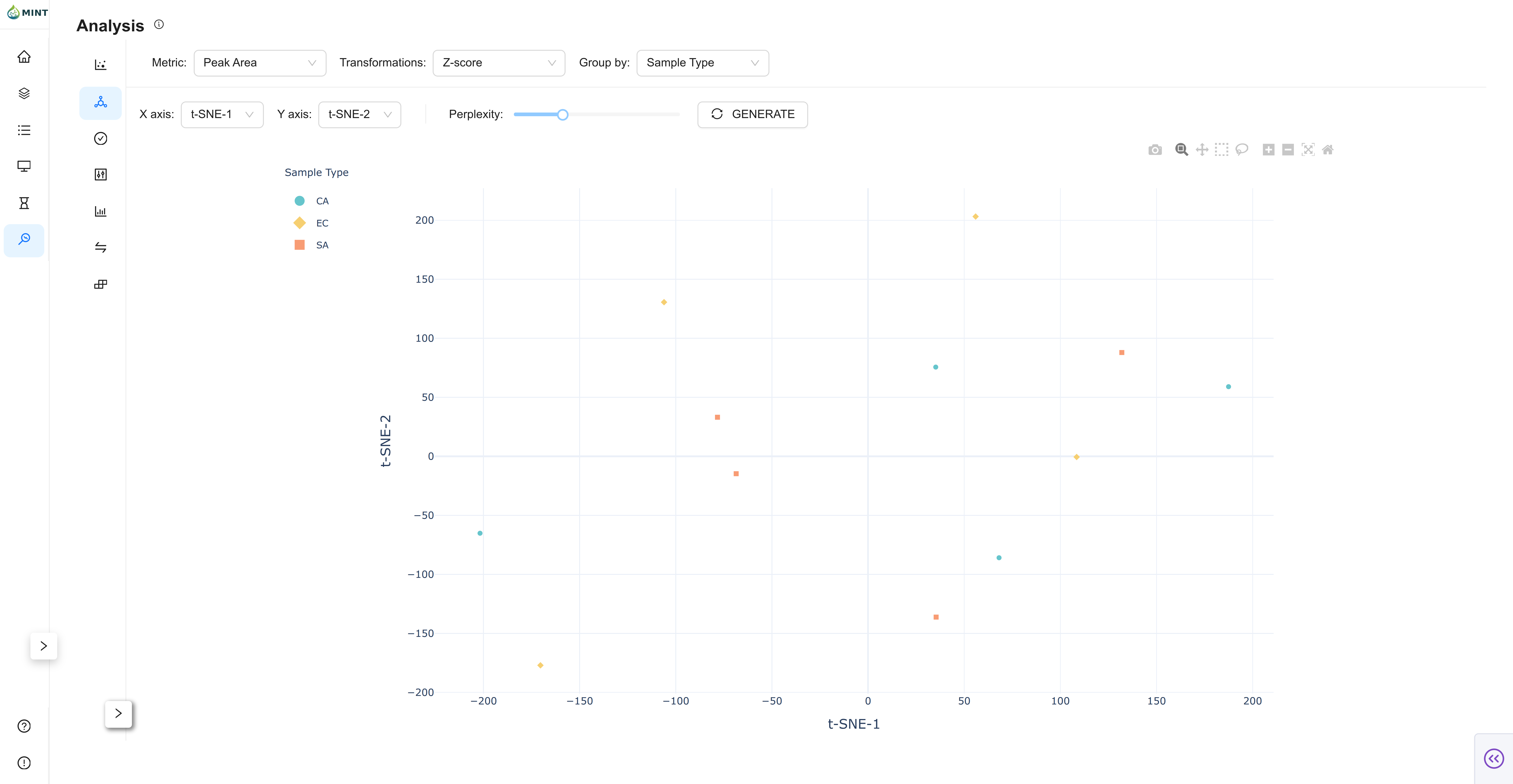
t-SNE View
t-SNE provides nonlinear embedding for local neighborhood structure.
- Axis controls: Choose displayed dimensions (
t-SNE-1...). - Perplexity slider: Tune neighborhood size.
- Generate button: Recompute embedding with current settings.
t-SNE computation details
- MINT computes up to 3 t-SNE dimensions.
- Perplexity is automatically constrained by sample count when needed.
Generateis useful when you explicitly want a refreshed embedding after parameter changes.
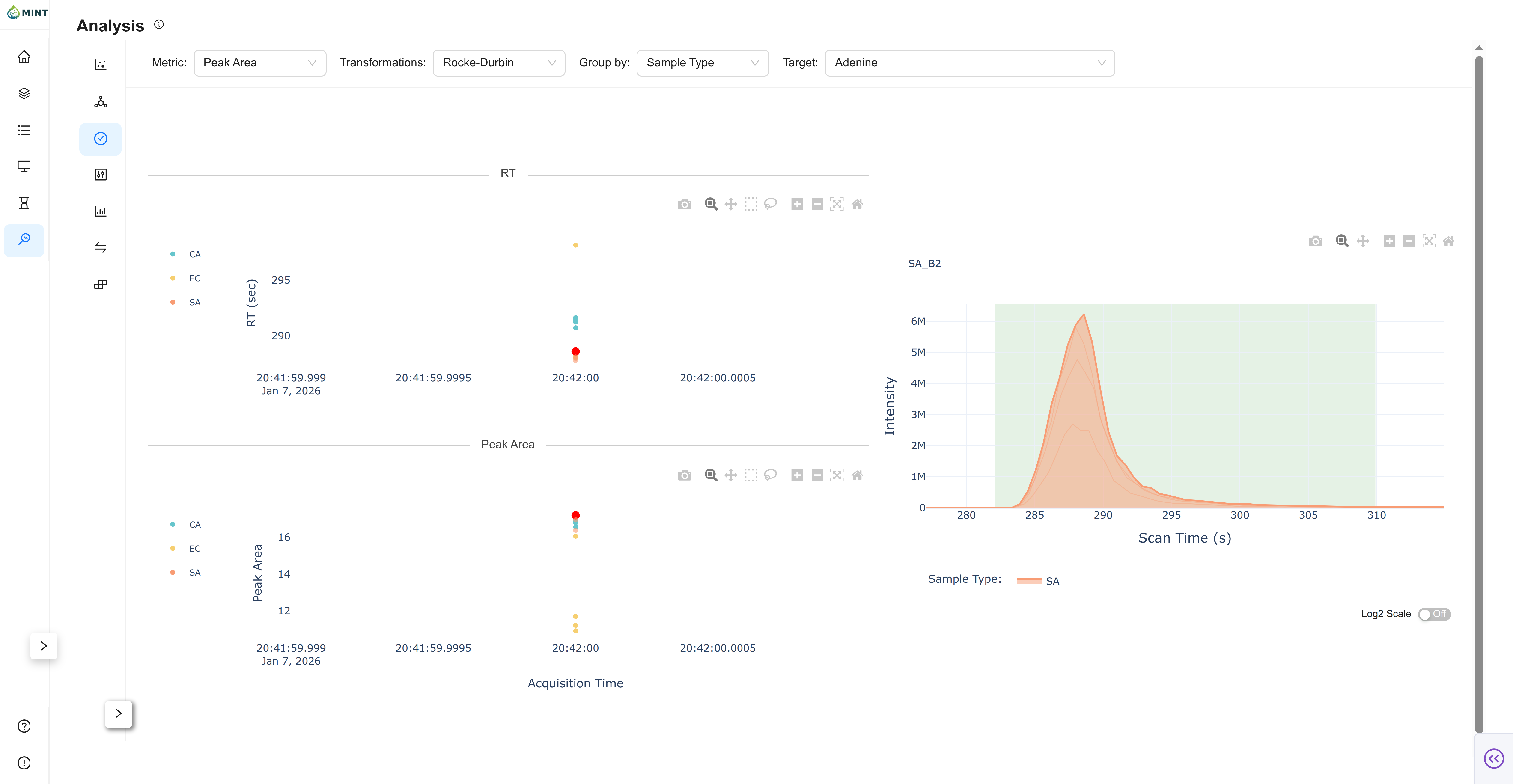
QC View
QC focuses on one target at a time.
- Target selector: Visible in the top bar only in QC mode.
- RT panel: Retention-time stability across samples.
- Metric panel: Selected metric across sample order.
- Chromatogram panel: Click points to inspect per-sample chromatograms.
QC target population and availability
- QC target options are populated from targets that already have results.
- Chromatogram behavior and grouping colors follow the same global
Group bysetting. - SCALiR concentration can be used as QC metric when concentration data exists.
- Large chromatogram views use Progressive Loading / Shadow Plots and LTTB downsampling to stay responsive.
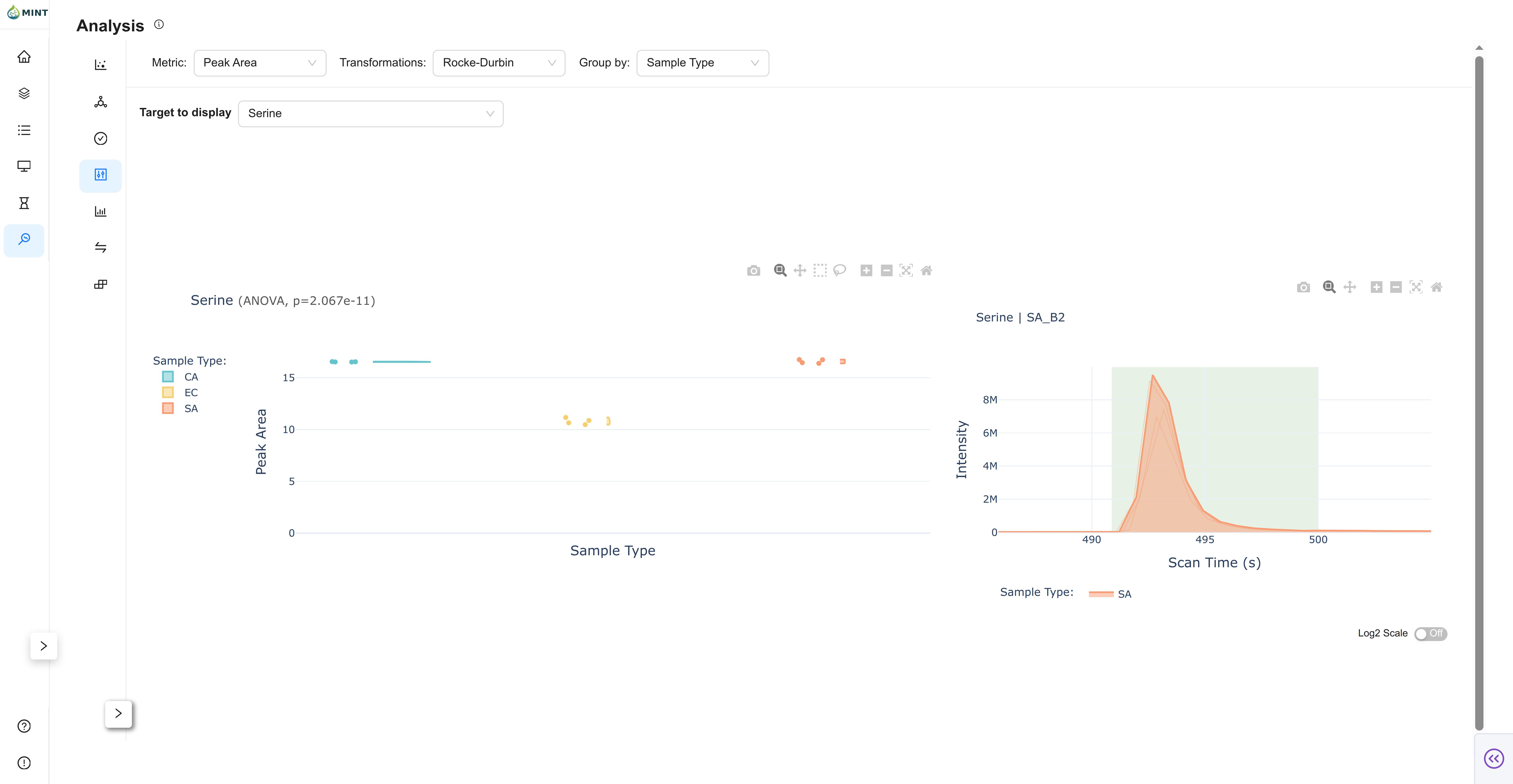
Violin View
Violin shows per-target distributions by group and links directly to chromatograms.
- Target selector: Search and select a compound.
- Distribution plot: Violin + points grouped by the selected metadata field.
- Stats in title: t-test (2 groups) or ANOVA (>2 groups), when applicable.
- Linked chromatogram: Click a point to open sample chromatogram.
Smart defaults in Violin
- Initial target defaults to a high-impact feature (PC1 loading heuristic).
- When needed, MINT auto-picks a representative sample with informative signal for chromatogram preview.
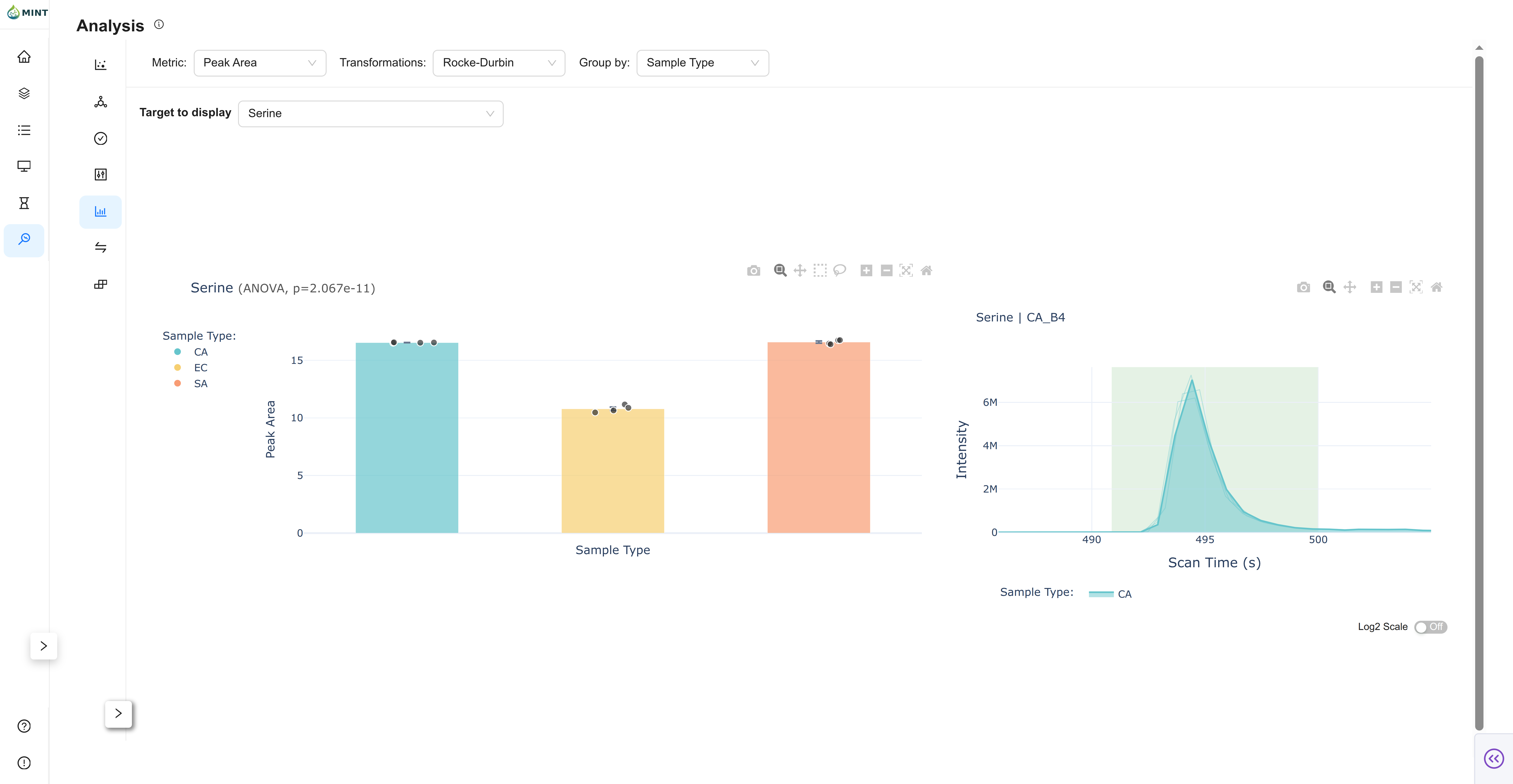
Bar View
Bar summarizes group means with sample-level context.
- Mean +/- SEM: Group bars with uncertainty.
- Individual points: Jittered sample overlay for spread inspection.
- Stats in title: t-test/ANOVA behavior mirrors Violin.
- Linked chromatogram: Click points to inspect sample traces.
Smart defaults in Bar
- Initial target defaults to a high-impact feature (PC1 loading heuristic).
- When needed, MINT auto-picks a representative sample with informative signal for chromatogram preview.
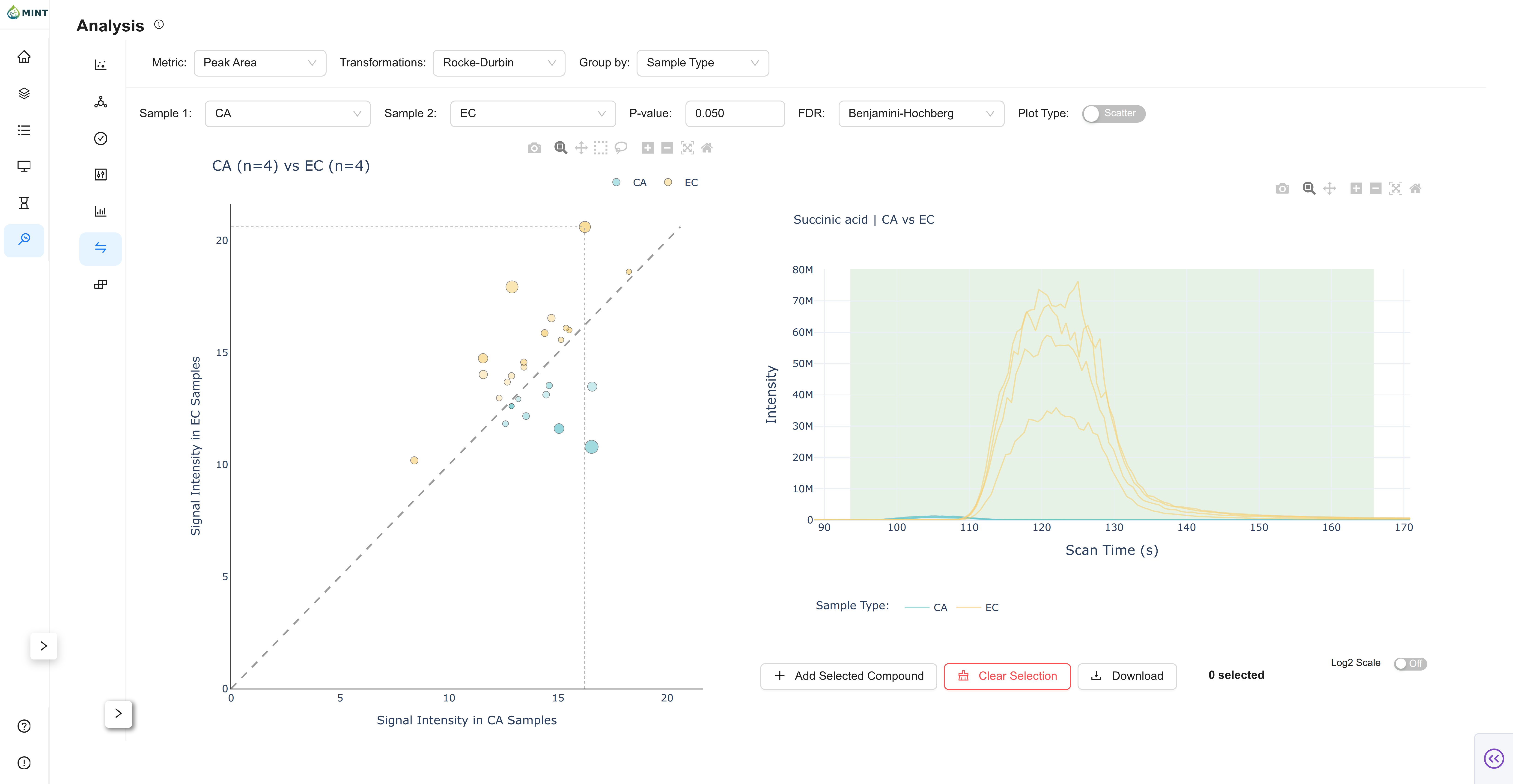
Comparison View
Comparison tests all targets between two selected groups.
- Group selection: Pick
Sample 1andSample 2from currentGroup by. - Statistics controls: Set p-value threshold and FDR method (
None,Benjamini-Hochberg,Bonferroni). - Plot mode: Toggle between scatter and volcano representations.
- Chromatogram linkage: Click a target point to inspect corresponding traces.
Feature selection workflow
Add Selected Compoundstores clicked features into a selection list.Clear Selectionresets the list.Downloadexports the selected feature list as CSV.- If the chosen grouping field has fewer than two distinct values, comparison inputs are disabled.
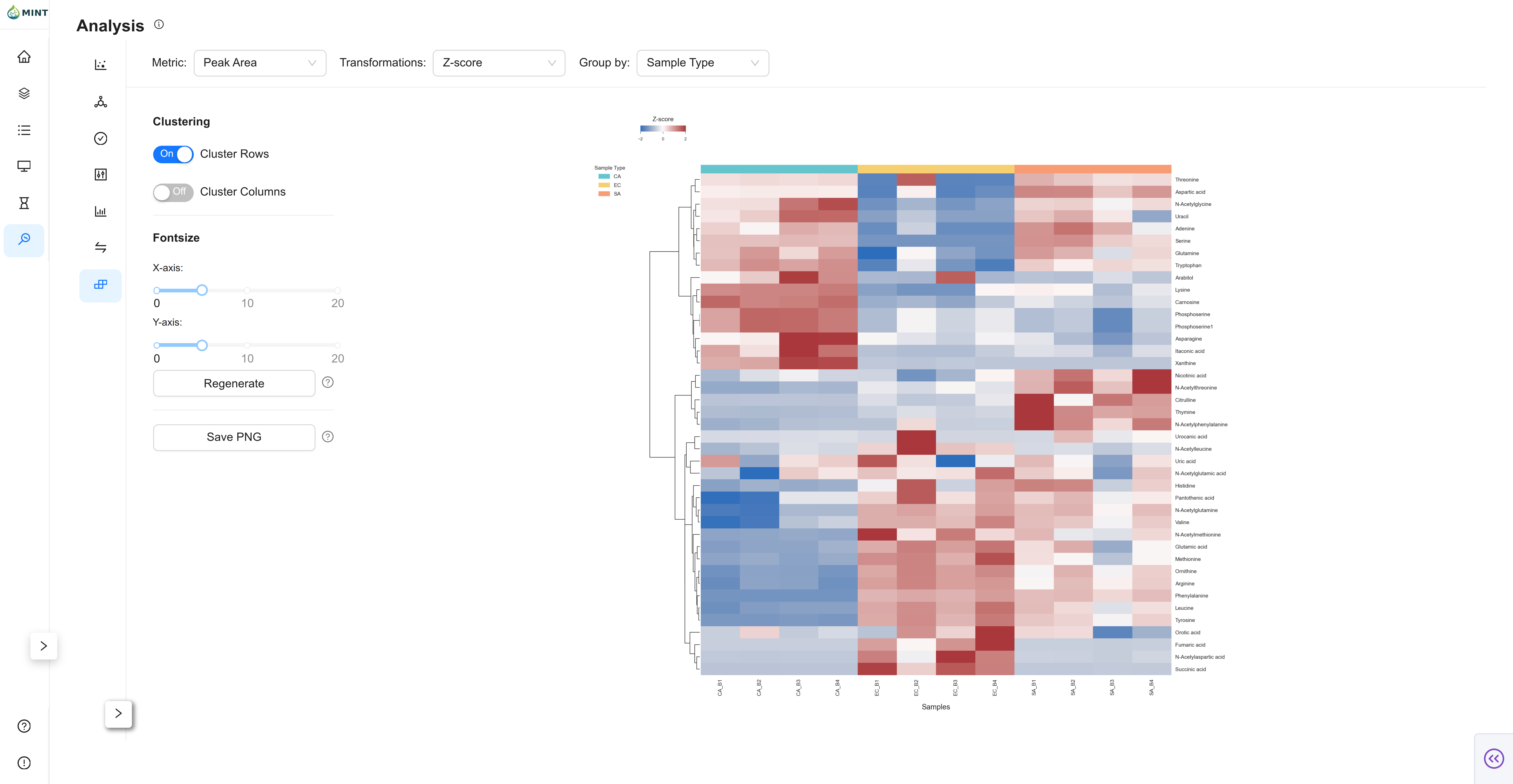
Clustermap View
Clustermap renders a hierarchical heatmap of samples and targets.
- Cluster controls: Toggle row and column clustering.
- Font-size controls: Tune X/Y label readability.
- Regenerate: Recompute the clustermap with current options.
- Save PNG: Download current clustermap image.
Clustermap persistence
- MINT also stores high-resolution clustermap images in workspace analysis folders for reuse/export.
Plot Downloads
All Plotly-based views can be exported as high resolution figures with standardized export filenames (date + workspace + view) from the modebar camera/download control on the top right corner of every plot, which helps maintain reproducible figure naming across sessions.
Performance niceties
- PCA/t-SNE/Violin include cache paths that reduce recomputation for group-only style changes.
- Spinners and staged updates are used across views to keep heavy updates responsive.
- Chromatogram-heavy views rely on Progressive Loading / Shadow Plots, LTTB downsampling, and sparsification.