Processing
The Processing tab is where MINT performs production extraction for your workspace, generating the final results table used by downstream analysis and exports.
Tip: Click the help icon (small "i" symbol) next to the "Processing" title to take a guided tour of this section.
Processing Workflow
- Set Scope and Resources: Click
RUN MINT. Choose bookmarked-only mode, recompute/fitting, and compute resources. - Run Extraction: MINT computes chromatograms, then results, then optional EMG fitting.
- Export Or Clean Up: Download results or remove selected/all result rows.
1. Set Scope and Resources
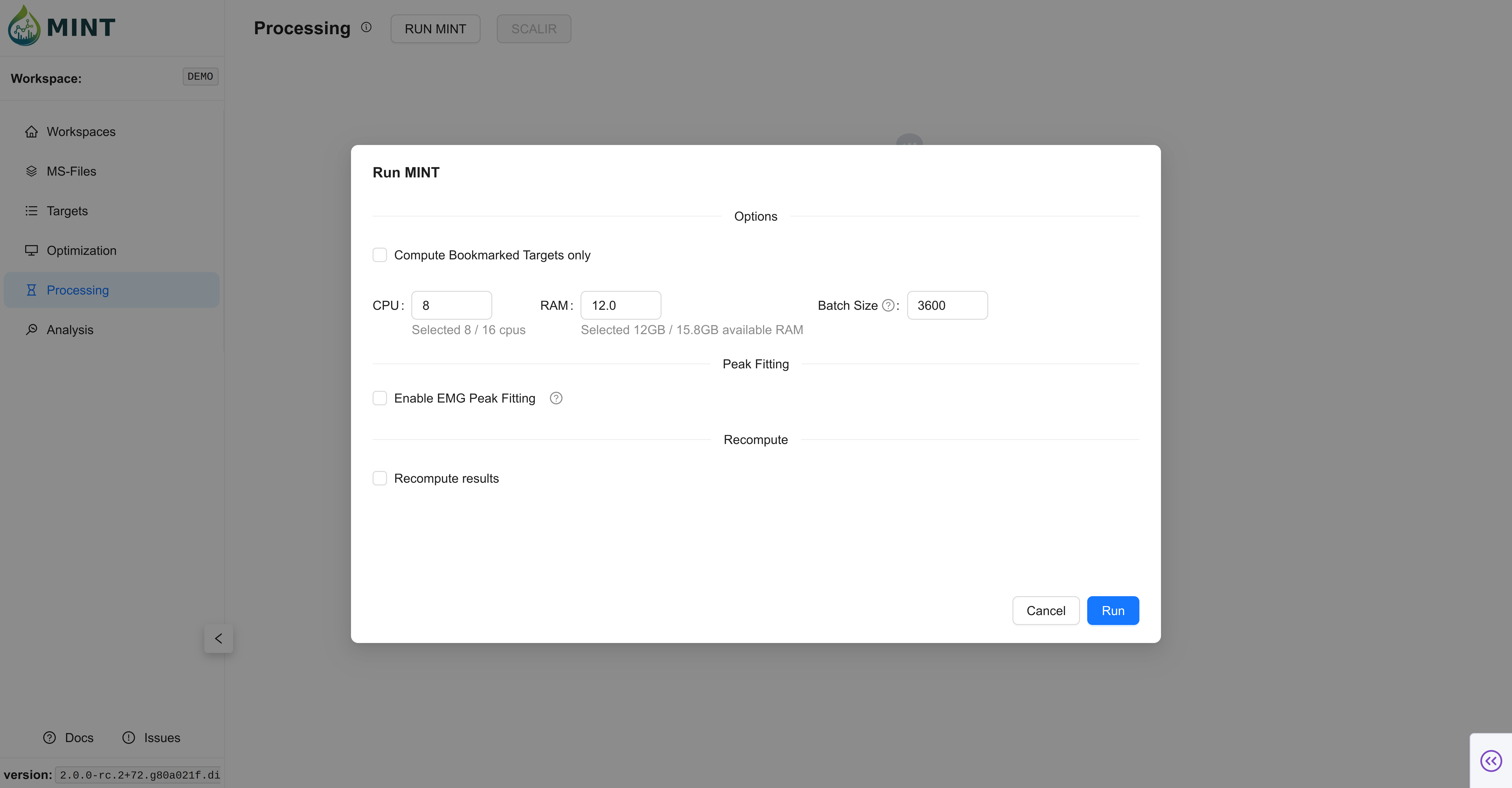
RUN MINT opens the execution modal for Processing.
- Bookmarked Targets Only: Restrict execution to bookmarked targets.
- Recompute results: Recalculate already existing result rows.
- Enable fitting (EMG): Run optional peak fitting after result extraction.
- Resources: Configure CPU, RAM, and batch size.
Run-time behavior and defaults
- CPU/RAM defaults are auto-detected from your machine.
- Batch size is auto-calculated from resource settings.
- Processing runs in staged phases with progress updates (
Chromatograms->Results-> optionalPeak Fitting).
What 'Recompute results' affects
- Recompute targets the results stage (and fitting stage, when enabled).
- During Processing runs, chromatogram generation uses the Processing pathway and is not the explicit MS1/MS2 recompute toggle used in Optimization.
2. Results Table
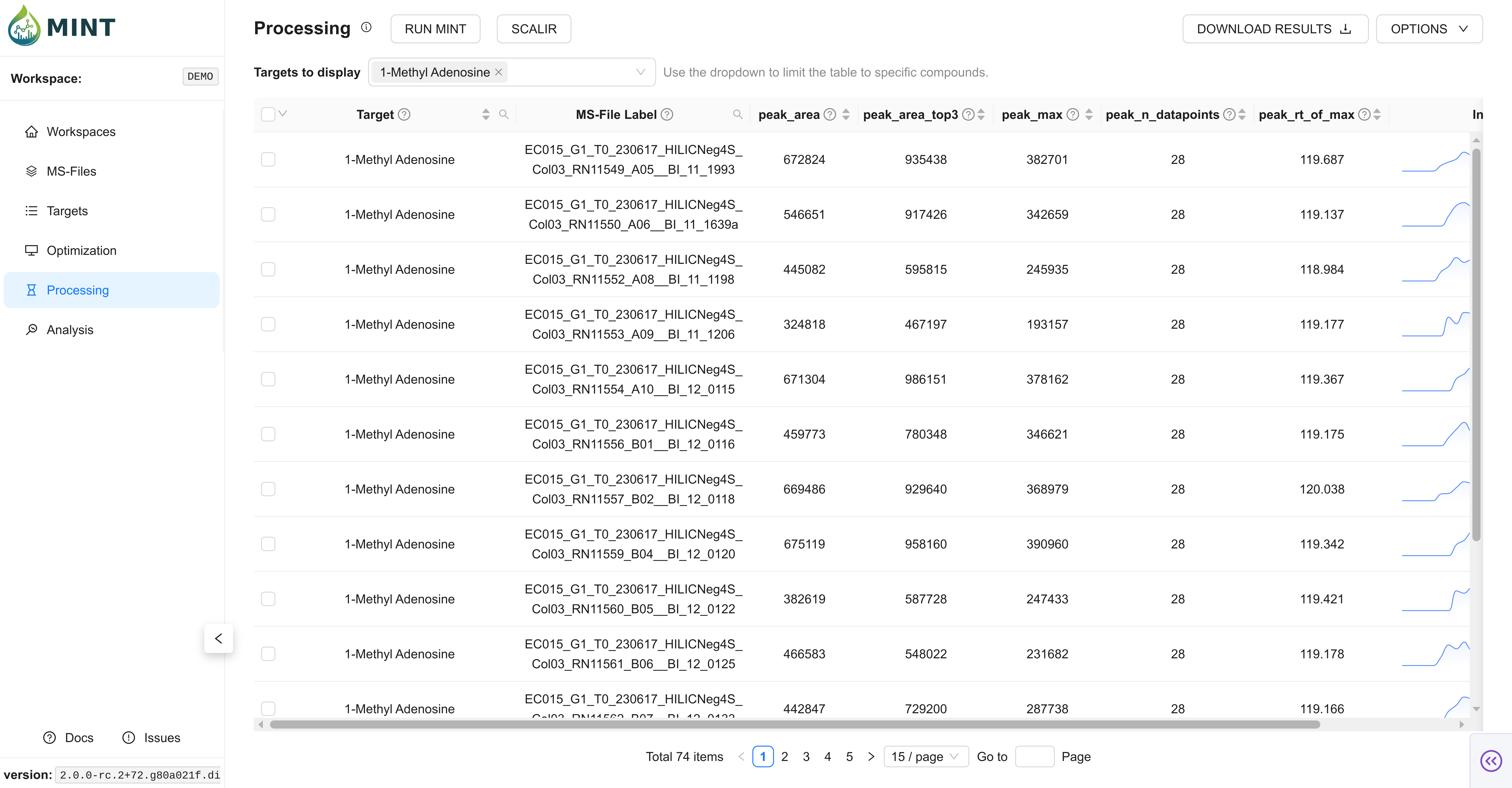
After completion, results becomes the primary working dataset for review and downstream steps.
- Target filter dropdown: Quickly narrow visible targets.
- Server-side filtering/sorting: Efficient browsing for larger result sets.
- Optional fitted metrics: Available when EMG fitting is enabled.
Core result columns
- Core extraction metrics such as
peak_area,peak_max,peak_mean,peak_rt_of_max. - RT-alignment fields such as
rt_aligned,rt_shift. - Optional EMG fit fields such as
peak_area_fitted,peak_rt_fitted,fit_r_squared,fit_success.
Processing filter semantics
- Results shown in this page are constrained to samples marked
use_for_processing = TRUE. - The target dropdown controls which compounds are rendered in the table.
3. Exporting Data
Use DOWNLOAD RESULTS to export either full (tidy/long) outputs or dense matrices.
- All Results (tidy/long): Column-selectable export of raw results rows.
- Dense Matrix (wide/pivot): Pivot by selected row/column/value fields.
All Results export behavior
- MINT validates selected columns against supported export fields.
- Export prefers a workspace backup file when available for faster delivery.
- For large files, MINT switches to direct Flask file download to avoid browser/base64 bottlenecks.
Dense matrix validation
- Requires valid row/column/value selections.
- Prevents invalid combinations (for example identical row and column keys).
Managing Results
Use the Options menu to clean up result subsets or reset the table.
- Delete selected results: Removes selected
(peak_label, ms_file_label)pairs. - Clear table: Removes all rows from
results.
Deletion semantics
Delete selectedremoves only selected target/sample pairs.Clear tableremoves the completeresultstable content.- Deletions invalidate the results backup snapshot so exports reflect current DB state.
Results Backup Snapshot
After a successful Processing run, MINT writes a workspace-local CSV backup snapshot:
- Path:
<workspace>/results/results_backup.csv - Purpose: Faster exports and resilience/recovery.
- Update behavior: Regenerated after runs; invalidated when results are deleted so the next export rebuilds cleanly.
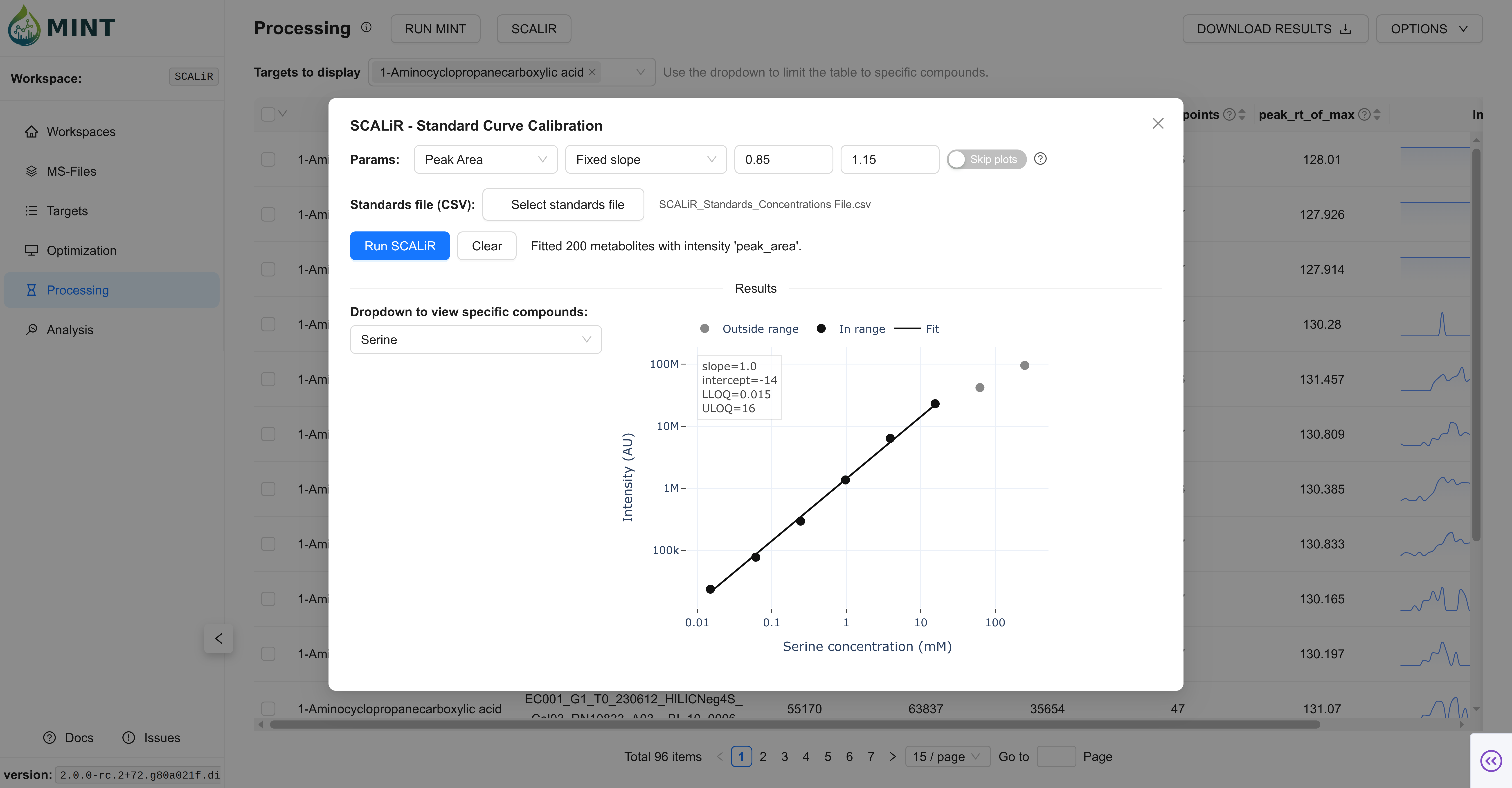
SCALiR (Optional)
SCALiR is an optional absolute-quantification workflow layered on top of Processing results.
- Provide a standards file.
- Fit calibration curves.
- Export concentrations and (optionally) calibration plots.